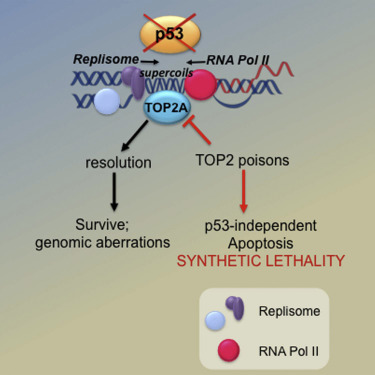
Molecular Vulnerabilities in Cancer
We identify molecular vulnerabilities in cancers by studying DNA damage response networks and cellular metabolism. Our work has uncovered non-canonical functions of the p53 tumour suppressor in safeguarding DNA replication, including how p53 prevents transcription-replication conflicts and how a p53-dependent NRF2-PARP1 axis links redox metabolism to genome integrity. Through high-throughput screens, we have identified metabolic vulnerabilities in p53-deficient cancers that can be exploited therapeutically.
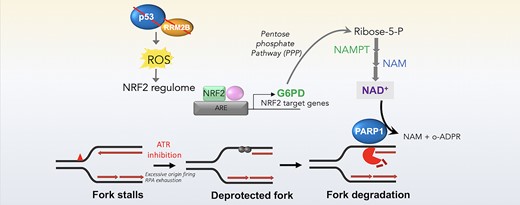
Graphical abstract -p53-dependent crosstalk between DNA replication integrity and redox metabolism via NRF2-PARP1 axis (Elfar et al., NAR 2024)
Key Publications
- Elfar et al., Nucleic Acids Research (2024) -NRF2-PARP1 axis
- Aning & Cheok, J. Mol. Cell Biol. (2019) -drugging in the absence of p53
- Kumar et al., Nature Communications (2018) -mitochondrial uncoupling in p53-defective cancers
- Kumar & Cheok, Scientific Reports (2017) -RIF1 SUMOylation and genomic stability
- Yeo et al., Cell Reports (2016) -transcription-replication conflicts